Candida auris: The path of yeast resistance
Dr Alexander Lorenz and his team are working to find out how the pathogen is able to colonise and infect humans and discover the key to its antifungal resistance.
Over ten years ago, C. auris was found for the first time in a Japanese patient’s ear canal. Today, the newly emerged pathogen is considered a leading cause of hospital-acquired fungal infections and one of the most drug-resistant yeasts known to man. Currently, more than 90% of isolates are resistant to fluconazole – the primary antifungal used to treat Candida infections – and many exhibit resistance to more than one of the three major antifungal classes. As a result, C. auris is difficult to treat and eradicate from intensive care units. Indeed, bloodstream infections have mortality rates of up to 50% and clinical alerts have been issued by US, EU and UK health authorities.
Despite its clinical significance, little is known about where the yeast came from or how it became capable of colonising and infecting humans. Researchers have yet to isolate C. auris outside a hospital, and the yeast has radiated from clinics in East Asia, South Asia, South Africa and South America to appear in the Middle East, Australia, North America, mainland Europe and the UK. Additionally, information on the fungus’ general biology and life cycle remains elusive, making the correct identification and treatment of the pathogen in clinics more difficult.

Now, University of Aberdeen researcher Dr Alexander Lorenz and his team are studying C. auris to see how genetic diversity among isolates drives virulence and drug-resistance.
Shuffling the (chromosomal) deck
Fungal pathogens such as Cryptococcus neoformans, Candida albicans and C. lusitaniae, a close relative of C. auris, often reconfigure their chromosomes to help them adapt to stressful environments. Apparently, C. auris does too. After testing 26 clinical isolates from the four geographic ‘epicentres’, Lorenz and his team discovered that even the most closely related strains had varying chromosome numbers, with East Asian strains having five to seven chromosomes and South Asian isolates having six or seven, despite sharing relatively uniform DNA sequences. The key to their disparity lies, at least partly, in the DNA’s stability – or rather, its instability. According to Lorenz, between one and four chromosomes harboured repeated copies of a gene called ribosomal DNA or rDNA which have proven volatile in other yeasts. In fact, these ‘rDNA repeats’ are often involved in gross chromosome rearrangements – leading to increased genetic diversity, altered gene expression and potential adaptations to challenging environments.
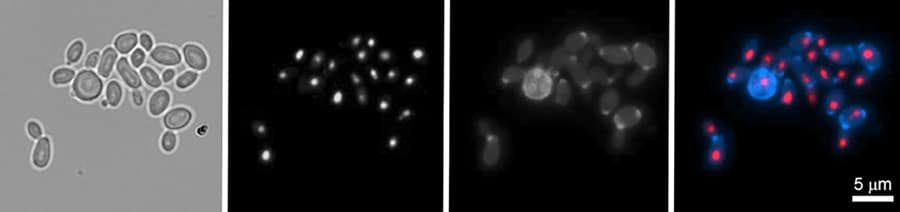
To see just how helpful these chromosome rearrangements can be, Lorenz and his team subjected four clinical strains (each from a different geographical hotspot) to heat, osmotic and genotoxic stress. To induce osmotic stress, the researchers exposed C. auris to a high concentration of the sugar sorbose. Osmotic stress is the sudden change in water movement across the cell wall (or cell membrane) caused by exposing cells to high concentrations of sugars or salts. Cells can adapt to this via stress response mechanisms. This process mimics the consequences of treating yeast with an echinocandin-type antifungal, which prevents cell wall synthesis. Meanwhile, genotoxic stress was achieved using a chemical that reduces the number of nucleotides available for DNA replication. Finally, heat stress was produced by growing the fungus at a sweltering 42°C.
Since [2009], the multidrug-resistant fungus has become a ‘major healthcare threat, with hospital outbreaks occurring worldwide’.
In all cases, chromosomes appeared, disappeared or changed sizes, with frequency and type of alteration differing depending on the strain and condition used. However very few modifications – mainly those in East Asian and South American strains – actually resulted in improved growth. It’s worth noting, though, that they did not appear to hinder viability or growth either.
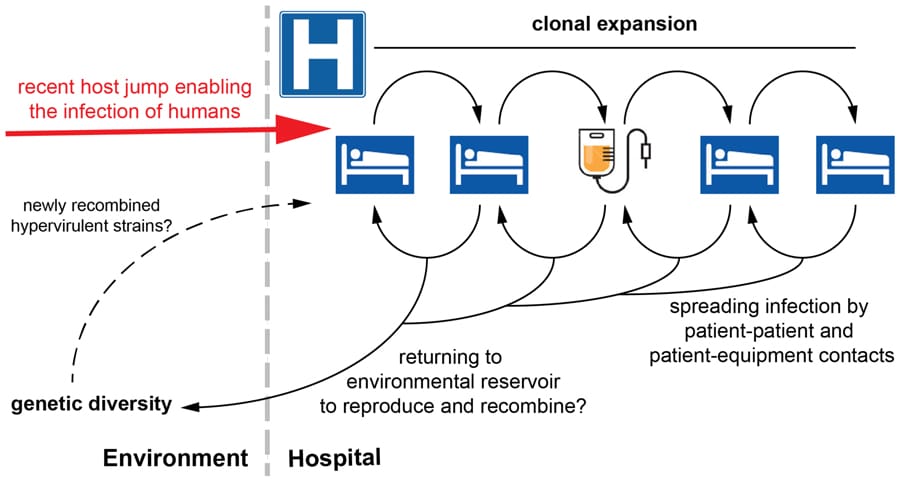
This is especially striking in the South African strain, which continued to grow after consolidating its genome from seven to three chromosomes when placed under heat stress. The yeast likely achieved this, either by tethering chromosomes from end-to-end or breaking its chromosomes apart and fusing the pieces to other chromosomes, according to the researchers. Such drastic changes could provide opportunities for adaptation rather than squander the cell viability of C. auris, the researchers added.
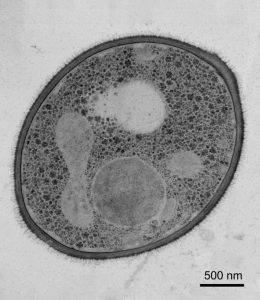
A selection of tricks
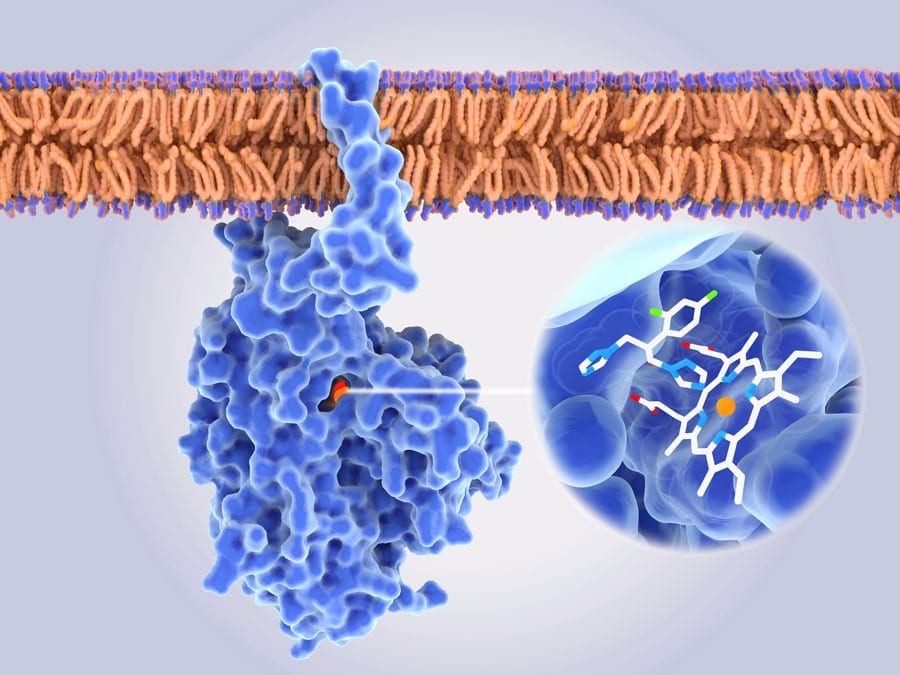
Chromosome shuffling isn’t the only adaptive trick up C. auris’ sleeve. The pathogenic yeast is studded with transporter proteins, capable of pumping antifungal drugs out of the cell. These proteins are more frequently expressed in the presence of antifungals such as azoles and polyenes, bolstering the yeast’s defences against the distressing compounds.
Additionally, under certain genotoxic conditions, many C. auris strains are capable of forming filaments called pseudohyphae. While there is insufficient data to confirm the role of pseudohyphae in the virulence of C. auris, filamentous growth has been shown to play an important part in host-pathogen attachment and disease development in other fungi. Combined with the malleable and adaptable genome of C. auris, this ability could allow clinical isolates to harness filamentous growth during pathogenesis.
… even the most closely related strains had varying chromosome numbers … despite sharing relatively uniform DNA sequences.
Researchers show their hand
While C. auris has shown itself to be a formidable and ‘mysterious’ organism, Lorenz and his team have found a potential way to fight back – at least in the clinic. Instead of an antifungal drug, however, the researchers used another fungus. When cultured together with C. auris, a predatory yeast, Saccharomycopsis schoenii, attacked and killed its cellmates using specialised lance-like structures. Within six hours, S. schoenii had eliminated 34% of C. auris, while almost no C. auris cells died when cultured on its own. Importantly, drug-resistant forms of C. auris were still susceptible to the predator. The results make S. schoenii a promising, if novel, candidate for biocontrol.
Next steps
In the 11 years since its discovery, C. auris has gained a reputation as one of the biggest causes of hospital-acquired fungal infections and one of the most drug-resistant yeasts known to science. Despite its global prevalence and dangerous nature, little is known about where the fungus came from, how it came to infect humans or how it developed antifungal resistance. To help fill these knowledge gaps, Lorenz and his team have been studying
C. auris – exploring its adaptability, genome structure, virulence factors and even ways to combat the fungus in clinics. Already, they’ve made considerable progress, but there’s still a lot left to learn about this mysterious organism, including its general biology, life cycle, additional contributors to its genome plasticity and how to defeat the yeast in patients.
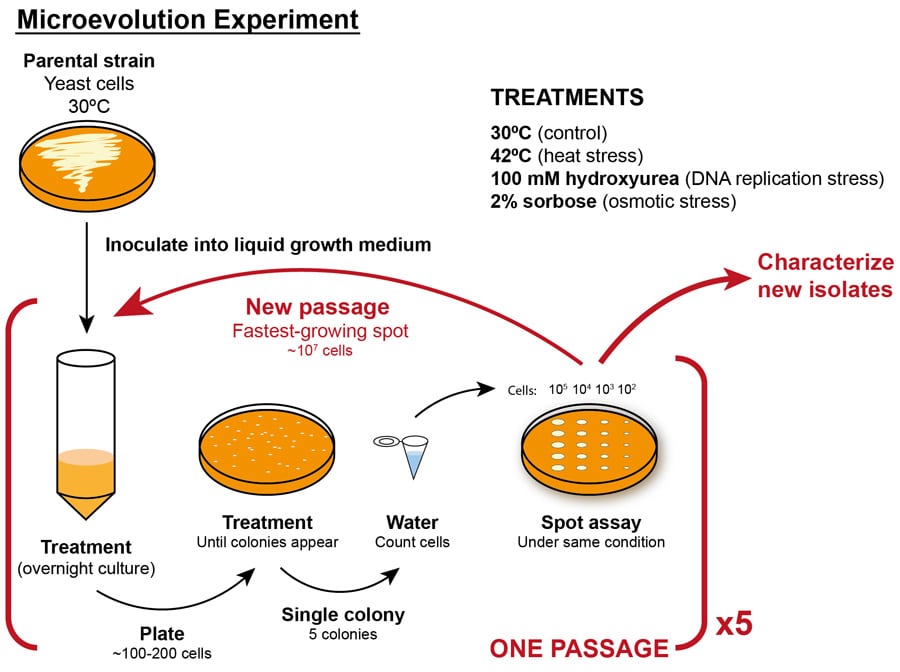
different geographical location, were subjected to heat, osmotic and genotoxic stress.
Personal Response
What prompted you to study C. auris?
Do you plan to continue studying the yeast, and if so, have you got any ideas for your next project?