‘Pool and split’ high-throughput experimentation: Turning complex designs into simple assays
Recent advances in technology and instrumentation have enabled scientists to increasingly benefit from high-throughput experimentation (HTE) protocols. These protocols allow scientists to perform multiple experiments and test multiple hypotheses, in parallel, and with minimal consumption of resources and materials. HTE protocols minimise reaction volumes, thanks to the use of micro-titre plates. These plates contain multiple wells – usually 96 – each acting as a miniature test tube, and each holding reaction volumes as small as a fraction of a millilitre. HTE is most advantageous in the fields of chemical synthesis or drug discovery, as multiple reaction conditions can be screened at the same time, increasing the probability of identifying useful synthetic pathways or active compounds.
Increasing the effectiveness of high-throughput experimentation
One disadvantage of HTE is that it requires intensive and laborious experimental design, as it involves the preparation of a very high number of different reaction mixtures. One further complication is that HTE intrinsically generates a very large number of raw data points calling for sophisticated data-capture and data analysis statistical software for the interpretation of results and data mining. For these and other reasons, the development of simplified approaches that can substantially reduce the experimental load and that do not need specialised instrumentation is highly desirable in the synthetic and pharmaceutical industries.

Dr Marco Santagostino and his collaborators, from the Chemical Process Development Department at Boehringer Ingelheim Pharma, Germany, devised a new protocol that greatly reduces the number of experiments needed in HTE. The team published a paper in 2021, applying their method for the quick identification of the optimal conditions for the cross-coupling reaction between carbon and nitrogen, when copper is used as a catalyst.
The authors named their approach ‘pool and split’, as it is based on a first ‘pooling’ stage, where complex mixtures of pooled compounds are screened for catalytic activity (i.e. increasing the rate of a reaction without being consumed), followed by a second ‘splitting’ step whereby gradually less complex mixtures are ‘split’ into their active components, so that the latter can be chemically identified and unambiguously characterised for their chemical properties and desired reactivity.
The development of simplified approaches that can substantially reduce the experimental load and that do not need specialised instrumentation is highly desirable.
The pooling stage: matching ligand sets to the perfect solvent
Dr Santagostino and his team devised a simplified HTE setup that would allow an easy identification of optimal conditions for copper-catalysed cross-coupling reactions for the formation of carbon-nitrogen bonds. To do this, they analysed arrays of catalytic mixtures of complex composition in 24-well plates. 24 copper ligands were pooled in four sets that were combined with mixtures of four copper sources – copper iodide, copper chloride, copper (I) oxide and copper (II) oxide – and three bases – potassium carbonate, potassium phosphate, and caesium acetate – in six mostly polar, high-boiling solvents.

Photo Credit: https://pubs.acs.org/doi/10.1021/acs.joc.0c02392
The ‘pool and split’ protocol allows 1728 reaction combinations to be evaluated in a single plate, allowing 24 parallel experiments to be followed at the same time. In each well of the plate a total of 72 combinations can be evaluated in one reaction, increasing the screening efficiency and reducing the analytical burden.
The first step of the ‘pool and split’ sequence consists of sorting the 24 copper ligands into 4 ligand sets, each set containing 6 ligands pooled together. Each ligand set meets specific criteria, such as the presence of specific chemical groups, similar reactivity patterns, or physical properties. The aim of this first screening step, performed in a 24-well microplate, is to select the optimal solvent to be used for each ligand set and identify the best ligand set. Specifically, ligand set 1 contained standard aliphatic nitrogen-containing diamines and ligand set 2 was mainly composed of oxygen-containing ligands such as polyols and di-carbonyls. Ligand sets 3 and 4 were composed of various oxygen- and nitrogen-containing aromatic compounds. The ligand sets were prepared by mixing the indicated ligands in equal amounts.
The deconvolution steps
The second step is also a 24-reaction run and involves the simultaneous ‘deconvolution’ of the most active ligand set and of the copper source mixtures in the solvent selected in the first discovery screening run. Deconvolution of the ligand set means reversing the process undertaken in the pooling stage by investigating the six ligands from the set, in isolation. As seen with the first step, the deconvolution phase is also conducted in a 24-well microplate.

The last step concludes the deconvolution stages of the ‘pool and split’ protocol, focusing on just one combination of ligand, copper source and solvent, exploring in three additional experiments the effect of choosing a base among the three available to test: potassium carbonate, potassium phosphate, and caesium acetate. This last step is normally conducted on a larger scale in a 10-well parallel reaction setup.
The ‘pool and split’ protocol allows 1728 reaction combinations to be evaluated in a single plate.
Dr Santagostino and his colleagues also developed an in-house user-friendly data analysis workflow. Visual Basic scripts were used to automatically extract data on the yield from HPLC chromatography outputs (high-performance liquid chromatography is a technique used to separate and identify components in a mixture). This allowed for visualisation through the TIBCO Spotfire software, for the purposes of data mining and graphical display in the form of pie charts. Each pie sector indicates the amount, in terms of experimental yield, of key reaction components (e.g., if decomposition reactions occur that produce other substances, besides the desired product, the output will be a total pie area below 100%). The method was validated on four cross-coupling reactions previously reported in the literature, in each case confirming the reported conditions. In all cases, improved conditions were identified by using the ‘pool and split’ approach that were not employed in the reported literature conditions.
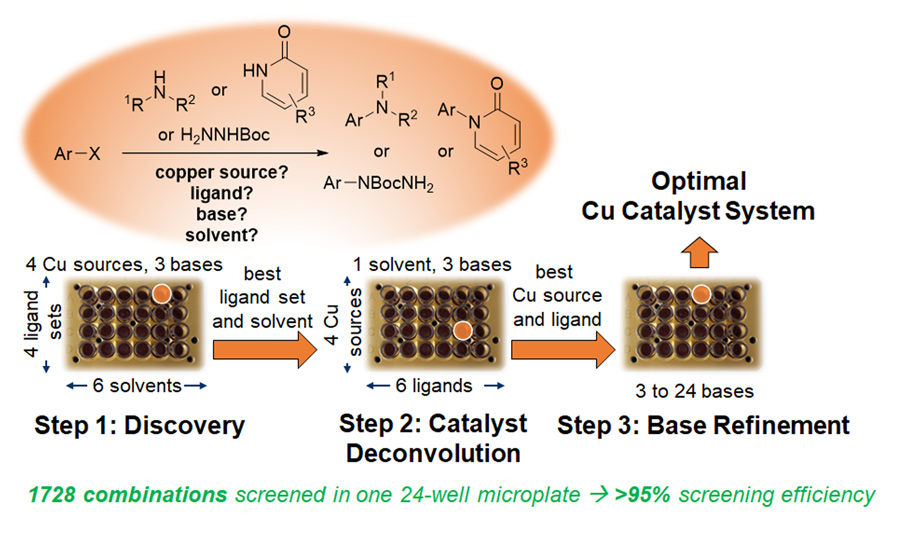
Photo Credit: https://pubs.acs.org/doi/10.1021/acs.joc.0c02392
Advantages of the technique
The researchers from Santagostino’s laboratory at Boehringer Ingelheim Pharma, Germany, and their collaborators, have developed the ’pool and split’ approach to simplify high-throughput experimentation. Despite following a high-throughput experimental design, the proposed method can be easily followed relying on inexpensive technological platforms, such as 24-well microplates. Key to the success and efficiency of this approach is also a user-friendly data analysis workflow. The researchers argue that the method offers a screening efficiency higher than 95%, which can be achieved with minimal costs. This approach has been applied to carbon-nitrogen cross-coupling reactions mediated by copper, a metal that is more sustainable as a catalyst than the rare and precious palladium. Carbon-nitrogen cross-coupling reactions are extremely important in the pharmaceutical industry because of the high content of nitrogen-containing molecules used in drug manufacturing.
The ‘pool and split’ approach facilitates the identification of optimum reaction conditions. This allows researchers to avoid large numbers of experiments typical of combinatorial approaches that would normally be needed in order to investigate thousands of possible combinations. The team has greatly benefitted first-hand from their optimised version of high-throughput experimentation, which has enabled them to design scaled up industrial synthetic steps for catalysis, supporting the large-scale preparation of new medicines. The method also simplifies the analysis and interpretation of the data and with further adaptations will enable the use of lab automation technologies. The research team anticipate that their strategy will enable the screening and optimisation of a very broad range of chemical transformations, proving to be a game-changer in the field of high-throughput experimentation.

Personal Response
What type of reactions do you plan to explore next with your ‘pool and split’ method?