New synthetic biology method revolutionises DNA cloning
Synthetic biology is a multidisciplinary field combining principles from molecular biology, chemistry, computer sciences, engineering, biophysics and others. At its core, it aims to engineer biology to help find creative solutions and developments in medicine (like new drugs and vaccines), biotechnology, bioenergy, novel materials, and the environment, to name a few. Synthetic biology allows scientists to better understand how the building blocks of life work and how to make use of them as machines. It is like a biology-based “toolkit” and its implications for research and development are endless. It allows the design and construction of new/modified biological devices and systems (cells/organisms) to synthesise novel molecules and alter existing biological systems to produce valuable, useful products. This could be construction of novel artificial enzymes (the catalysts of cells) or biochemical pathways and genetic regulatory circuits. Currently, synthetic systems are already used to mass-produce medical compounds like insulin and cancer drugs.

Synthetic biology is comprised of four principles. The first is that if we truly understand a biological system then we should be able to reconstitute it from its basic parts. Second, if we manipulate these systems, we can understand them and the basic principles of life better, as well as having the ability to manipulate and create novel systems. The third principle is that living systems have not been optimised for human understanding but have instead “evolved to continue to exist.” So, by engineering living systems it is possible to simultaneously test our current knowledge and create new models that are easier to study. For example, in 2016 researchers designed a synthetic organism with a minimal genome – a genome reduced to only the essential genes required for life. The fourth, and final principle, is the idea that biology can be deployed as a technology.
Advances in synthetic biology have been made possible by developments in DNA sequencing and DNA synthesis. These two technologies have allowed scientists to read and synthesise DNA less expensively and more rapidly than ever before. To date, however, a third enabling technology, molecular cloning, has not kept pace with technological advances made in DNA synthesis and DNA sequencing.
Generally speaking, in order to create devices and systems, each individual part (a DNA sequence possessing a defined function) must be inserted into an individual plasmid, amplified and isolated before it can be combined with additional parts to make higher order devices. Then, by stitching together multiple devices which produce one or more proteins and inserting them into the same cell you can create a system, such as a biosynthetic pathway.
Synthetic biology has allowed scientists to better understand how the building blocks of life work and how to make
use of them as machines.
Plasmids can contain a variety of informative parts, such as one or more genes coding for the synthesis of scientifically relevant proteins or mutated versions of a gene. All plasmids contain some essential parts, such as an origin of replication (OR), which directs bacteria where to start duplicating it. Nearly all plasmids include a selection marker (SM), often a gene that confers resistance to a specific antibiotic, allowing only the bacteria that contain the plasmid to survive. Other parts include: promoters (P), a DNA sequence placed before a gene of interest (GOI) that instructs cells when and where to produce the GOI; ribosome binding sites (RBS), defining where ribosomes attach to RNA (ribosomes are cellular complexes that read RNA and translate it into proteins); stop signals (that tell the ribosome when to stop translating); and a variety of pre-(N) and post-(C) terminal protein tags.

EDITOR: Ph.D. Jeffrey Carl Braman, DOI: 10.1007/978-1-4939-7795-6, ISBN: 9781493977949 • 9781493977956 (online),
SERIES: Methods in Molecular Biology, Volume 1772, Springer, New York. This detailed book explores a few of the plethora of techniques and applications associated with the unique branch of science known as synthetic biology. Chemists, biologists, and engineers engaged in this multidisciplinary field of study will be guided in the creation and regulation of gene circuits, manipulation of biochemical pathways, genome editing and modification, creating genome language and computing, as well as molecular assembly. Written for the highly successful Methods in Molecular Biology series, chapters include introductions to their respective topics, lists of the necessary materials and reagents, step-by-step and readily reproducible protocols, plus troubleshooting tips and explanations of how to avoid pitfalls, all written by experts who have published their work in peer-reviewed journals. Authoritative and practical, Synthetic Biology: Methods and Protocols provides key guidance and ideas for conducting your own synthetic biology projects.
OR, SM, GOI and P are the basic components or parts that can be used in a plasmid. Once several thousand plasmid copies have been made, researchers are able to extract the plasmid molecules and transfer them into their cells of choice, be it cultured cells, embryos or whole organisms. Imagine the bacteria as a factory producing plasmid copies and the other recipient cells as processors of the plasmids.
Traditional and several recently developed cloning methods rely on the use of a variety of restriction enzymes to obtain desired parts and devices for creating systems in plasmids. Restriction enzymes (REs) recognise specific DNA sequences and cut them; a bit like creating a DNA puzzle using scissors where each scissor cut creates a distinct shape, so that the resulting DNA pieces can only be put together with complementary shapes. In conventional molecular cloning the DNA fragment (part) is isolated from the pieces that were just cut, and then ligated (glued) into a plasmid with a compatible end. This means that in order to create devices, the experimental design must take RE DNA recognition sequences into consideration. These recognition sequences must not be present within the part of interest as the part would also be cut. Therefore, the choice of REs is a major consideration of the design process.
SureVector represents a set of synthetic biology tools to quickly construct new combinatorial devices and systems allowing the research community to design products faster, less expensively and more efficiently than ever before.
This process requires hierarchical binary assembly, i.e. parts required to create a device are added together sequentially. This makes the whole process time consuming and laborious, resulting in very difficult combinatorial experimental design.

Here comes the revolution – SureVector
As previously mentioned, conventional molecular cloning methods rely heavily on the use of plasmids and REs. Jeff Braman and Peter Sheffield from Agilent Technologies developed a new method to assemble DNA parts that allows faster and combinatorial creation of devices and systems. They generated a host of parts for creating plasmids allowing expression of proteins in E. coli, yeast and mammalian cells. These parts each contain unique 30 base-pair (bp) DNA sequences to flank individual parts for ease of device and, eventually, system assembly. These sequences are unique in that they are not found in any of the DNA databases available online. By having the same 30 bp at the end of one part and the beginning of another, parts are seamlessly assembled in the desired order (Figure 1).
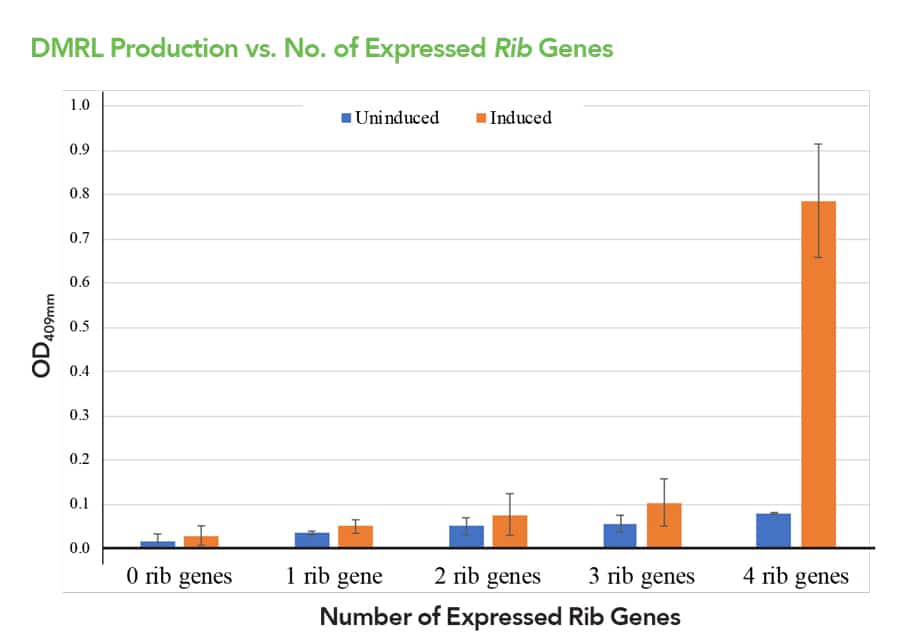
Compared to other current methods that convert parts into devices, SureVector does not require manipulation of parts with REs. Instead, all desired parts are mixed together, and due to the uniqueness of the flanking 30 bp DNA sequences they are assembled in the correct order using a specialised enzyme blend. One of the advantages of this new method is the ability to assemble multiple parts in a parallel fashion that “substantially simplifies and speeds optimisation of device and system development.” When designing a new system, the order and arrangement of the different parts and devices has implications for the efficiency of the system. This was demonstrated by Dr Braman and Dr Sheffield when they reconstructed the four-enzyme biosynthetic pathway for 6,7-dimethyl-ribityllumazine (DMRL), the precursor for vitamin B2. They were able to create in parallel a collection of devices with different combinations of parts that they then used to generate systems. These systems were tested for their efficiency to produce DMRL. In under a week, Braman and Sheffield were able to identify several combinations of devices leading to the creation of the best DMRL producing system (Figure 2).

Personal Response
How do you think this new technique will be helpful for researchers when compared with other techniques like Gibson assembly?
Other methods employ one or more enzymes to generate parts with single strand overhangs for the assembly process to be efficacious. These methods require parts to be at least 250 base pairs in length to prevent the enzymes from completely destroying the desired part.
Your technique massively speeds up and simplifies the assembly of synthetic biological systems. Where would you most like to see it being applied and why?
J. Braman: SureVector technology expands the domain of organic chemical synthesis by facilitating the ability to tailor design and speed synthesis and testing of fine chemicals, intermediates, and novel drugs employing combinations of unique enzymes and biosynthetic pathways.