Thermodynamic modelling of dynamic cell phenotypes
Cell-based therapies are an exciting branch of modern medicine. By modifying certain cells taken from a patient or donor, researchers have recently discovered new ways to treat diseases, ranging from neurological conditions to certain forms of cancer. Currently, however, many cell-based treatments remain in their early stages and must overcome several key challenges before becoming widespread in practice. To gauge how populations of cells will respond as therapies, developers and manufacturers must determine their ‘phenotypes’ or critical quality attributes, that describe the sets of measurable traits which are expressed by their underlying genetic code.
A major difficulty in doing this lies in the inherent unpredictability of the changes in phenotypes that can occur in individual cells over time, even within apparently stable populations. Since the traits of each cell are the result of many complex cellular reactions, researchers must rely on mathematical descriptions of populations of cells. “While a population of cells can show a highly stable range of phenotypes, individual cells within the population are highly variable and dynamic in phenotypic expression”, NIST Fellow Dr Anne Plant explains. “This knowledge allows us to develop mathematical expressions that describe the role of cellular fluctuations.” In their research, Dr Plant and her colleagues explore how these cellular characteristics can be described with mathematics that is typically used in a completely different field of science.
Applying statistical thermodynamics to cellular dynamics
Statistical thermodynamics is a branch of physics which describes the relationship between heat, mechanical work, and the flow of energy within physical systems. At first glance, the field may appear to be of little relevance to biological networks, but the diverse and dynamic nature of individual phenotypes within large, stable cell populations displays a remarkable similarity to the varying physical properties of individual gas molecules in a confined space. To describe these variations, physicists use mathematics first set out by Austrian physicist Ludwig Boltzmann in the 1870s.

According to Boltzmann, the properties of thermodynamic systems relate to the collective properties of large ensembles of particles. Crucially, a given state can result from any one of a vast yet finite range of ‘microstates’ which describe the unique positions and velocities of each particle in the system. However, depending on its temperature and energy, a system will not have an even probability of existing in all microstates across this range. Instead, this probability is defined by a distribution curve or its negative log which is a potential landscape, featuring a peak or valley respectively, at the microstates where the system is most likely to find itself. As biologists later discovered, clear connections can be drawn between the probability distributions of nonequilibrium thermodynamic systems, and those of phenotypes within large populations of cells.
Connections can be drawn between the probability distributions of nonequilibrium thermodynamic systems, and those of phenotypes within populations of cells.
Measuring complexity and dynamics of gene expression
Cell phenotypes are far more complex than the positions and velocities of non-living particles. Within individual cells, the expression of genes that control cell phenotypes arise from intersecting networks of different factors, including the rate of synthesis of information-carrying RNA and reactions involving intricate interactions between protein molecules. Because of this complexity, it is virtually impossible for researchers to quantify all of the details involved in how phenotypes of individual cells come to be, and why individuals are different from one another. Therefore, the NIST team focuses on identifying the most important measurable quantities by essentially folding all the minute, hard-to-know details of the underlying processes into the overall observations that they can directly measure.
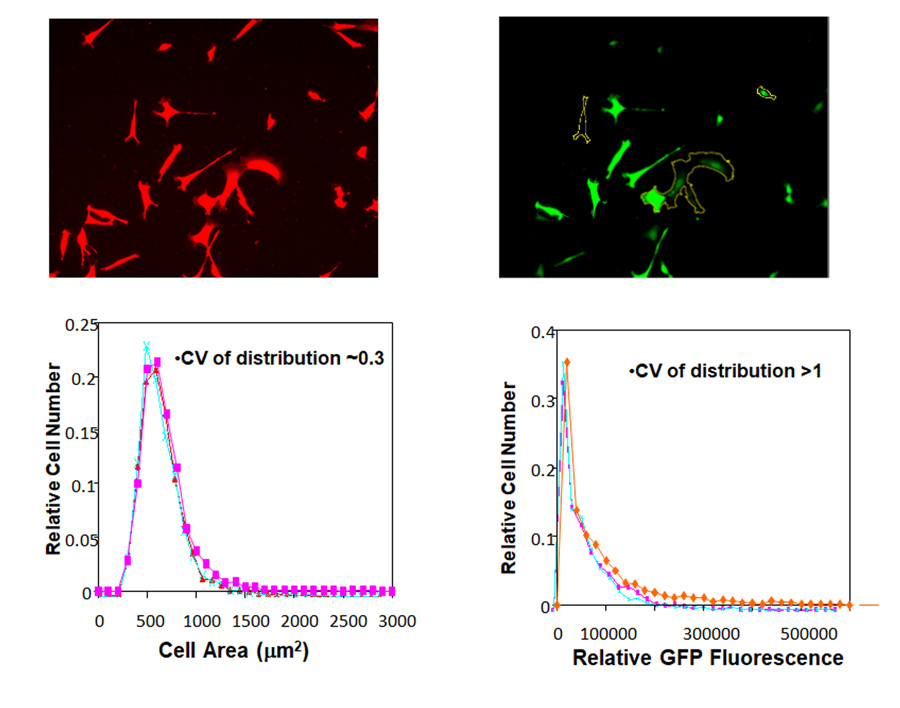
From these results, the researchers can gain important clues about the characteristics of sub-populations within larger groups of cells. These sub-populations, or microstates, contain cells with nearly identical phenotypes. “We focus on what can be experimentally measured, such as the rates of fluctuation of light-emitting protein reporters of genes of interest”, Dr Plant describes. “These provide mathematical quantities that allow us to predict how fast cells exit from a microstate to populate other microstates within the population.”
To visualise this concept, we can think of a landscape where sub-populations exist in valleys with varying depths, also called attractor basins. Moreover, the cells’ phenotypes are continually changing – they are like jostling marbles, constantly trying to escape from their attractor basins. In this analogy, deeper valleys correspond to microstates with more stable traits. As a result, cells will find it harder to escape from the sub-populations in attractor basins, meaning more cells will collect inside them. From here, Boltzmann’s probability distribution becomes relevant once again, with the more thermodynamically stable microstates being the ones containing more cells.
Entropy and relative free energy
As the team shows, this behaviour is directly comparable to two key aspects of thermodynamics. The first of these is entropy – a term generally used to describe the level of disorder in a system. In this context, it relates to the range of microstates that describe the macroscale properties of a certain gas: if the value is higher, the gas particles must be in a more disordered state. The second is the relative free energy, which is the amount of energy required to keep a system in a steady state, even when particles are constantly moving between different microstates.
In their latest research, the team have drawn direct parallels between each of these quantities, and stable, or ‘homeostatic’, cell populations – enabling accurate mathematical descriptions of their behaviours. “We have shown a direct connection between entropy, relative free energy, and the population of steady-state of phenotypes using Boltzmann’s mathematics”, says Dr Plant. “This analysis allows us to consider the energy required to maintain the biological network in its dynamic, homeostatic state.”
This may improve procedures including creating T-cells for fighting cancer cells.
A small number of master regulator genes
Graph models are commonly used by systems biologists to map out connections within networks of systems. By modelling the components of a regulated cellular network as ‘nodes,’ and their connections as ‘edges’, they allow researchers to describe how changes within one node will drive subsequent changes in the rest of the system connected to it. Based on their new understanding of phenotype dynamics, the team constructed such a model as a basis for computer simulations. With this model, the researchers could then explore which components must be measured to accurately determine the relative free energy of a real system of cells.
“Simulations revealed that when there are a limited number of genes, or master regulators, that control the network, the distribution of phenotypic responses will be highly asymmetrical”, Dr Plant describes. “This suggests that the shape of the steady state distribution of phenotypes in a cell population provides insight into how many network components need to be measured to define the state of the network.” Techniques that measure many genes may be providing a lot of information that is not all that important to the control of the network. The team’s new insight now provides an approach that can drastically cut down on the numbers of measurements required to gain the same amount of information. This discovery could have significant implications for future applications of cell populations.
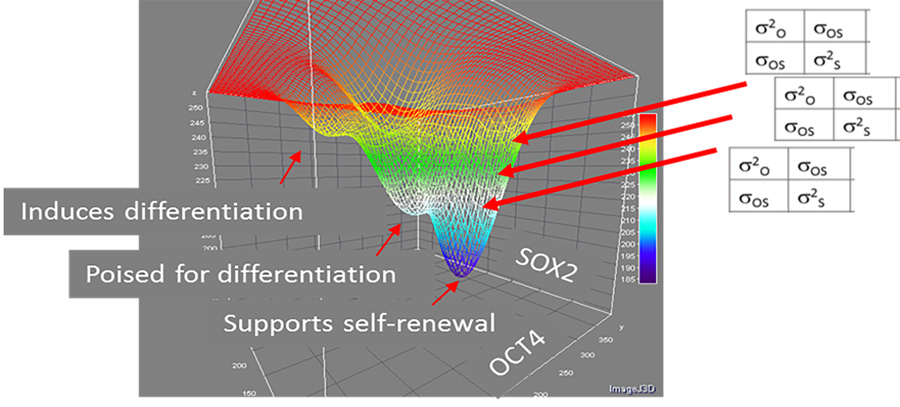
relationship between genes OCT4 and SOX2 over a population of pluripotent cells.
Improving cell-based manufacturing
The cells contained in our bodies have the remarkable ability to cure diseases inflicting other parts of the body. The technology required to create healthy cells to cure disease is now growing increasingly advanced – and through the NIST team’s discoveries could soon become yet more sophisticated. With a greater knowledge of the dynamic phenotypes of cells in populations, manufacturers and clinicians will be able to produce and deploy cell therapies far more safely and efficiently.
In turn, this may improve procedures including creating T-cells for fighting cancer cells; surgically applying stem cell-based healthy tissues for repairing damaged tissues; and potentially even renewing neurons in the brain to combat degenerative conditions such as Parkinson’s disease. The team continues to explore how their techniques can be advanced even further – bringing these exciting medical applications a step closer to reality.
Personal Response
How readily could your simulations be applied in a real cell-based manufacturing context, such as production of an immune cell therapy?