When the body turns against itself: Amyloid-based diseases
Proteins are large, complex molecules that are essential for making our bodies work. There are thousands of different kinds of protein, and each one has a specialist role. Some proteins help to drive out invading species like bacteria and viruses and form the basis of our immune systems. Other proteins help with the chemistry in our bodies, for example, helping to regulate the acidity of our blood and control carbon dioxide levels. Some membrane proteins even control electrical currents in our nerves and muscles.
What makes one protein different from another? Each protein has a unique structure, not just in terms of which atoms make up the chemical composition of the protein but its overall spatial arrangement. For proteins, their structure is often intimately related to their function and any deformities in their shape can hinder their work.
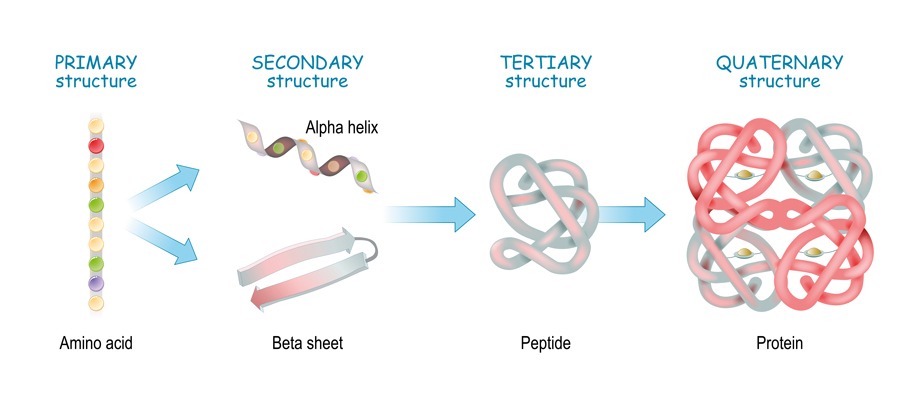
Protein structures are often classified into four levels. At the lowest level, you have the primary structure which describes the chemical motifs that make up the protein. These are amino acid chains, known as polypeptide chains, that are linked together by chemical bonds. Then, you have the secondary structure. The amino acid chains that make up a protein might contain over 500 amino acids, and so they often curl up and form complex shapes due to chemical interactions between different parts of the chain and the resulting structure is known as the secondary structure.
This predictability is an essential part of future drug design.
The tertiary structure of a protein is essentially its three-dimensional arrangement in space. Unlike the secondary structure, which describes localised structures in the overall protein, the tertiary structure takes into account the overall complex picture of its full shape. All proteins have tertiary structures, but some proteins have multiple polypeptide chains that act as subunits within the full structure of the protein. This gives an additional layer of structure, known as the quaternary structure and these subunits can sometimes behave independently of other regions of the protein to allow for more complex functionality.
The complexity of all of these layers of structure and the intricacy involved in shaping hundreds of chemical structures means that sometimes things go wrong, something that Dr Robert Guy at Amyloid Research Consultants (ARC) has been deeply interested in. He has been trying to understand how the shapes of proteins might relate to their role in diseases such as Parkinson’s and Alzheimer’s with the hope of finding a way to treat them.

Structure mishaps
Many devastating diseases including Alzheimer’s, Parkinson’s, type 2 diabetes, Creutzfeldt-Jacob (‘Mad Cow’) disease, and some cancers are related to the presence of amyloid proteins. Most are incurable or difficult to treat. Amyloids usually clump together to form large waxy, fibrous structures. These fibrils are easy to identify, and portions of their three-dimensional structures have been determined experimentally.
But it is becoming increasingly clear that fibrils are not the only, or maybe even the major, cause of these diseases. What makes these amyloid-based diseases so difficult to cure is a quirk of the amyloid proteins themselves. Before forming fibrils, amyloid proteins form a large range of smaller aggregated structures, called oligomers and protofibrils. The number of polypeptide chains within oligomers can vary from two to hundreds. As their size increases, they adopt a hugely diverse range of secondary structures. Some have sheet-like assemblies, some helices, and some are disordered.

Knowing a protein’s structure and functional mechanisms are key to finding effective drugs and antibodies for treating protein-based diseases as the shape of the protein affects which chemicals can bind and alter the protein’s effects. Thus, despite the challenges of trying to obtain it, many researchers have performed experiments and computational modelling to try and solve unknown protein structures.
Dr Guy has also been investigating an additional dimension of complexity to the protein structure problem – dynamical changes. The number of polypeptides comprising amyloids increases with time from only a few to thousands of fibrils. As they grow and spread to different cellular environments, all levels of their structures change dramatically. Some assemblies adopt structures vital for normal brain functions, but others deviate to toxic structures. Electrical impulses both within and between neurons are caused by the transient opening of proteins, called channels, that span the neuron’s membrane. Most amyloids not only alter properties of some channels, they also form channels by themselves. Altering or ‘short-circuiting’ the brain’s normal electrical currents is the last thing one would want to happen.
Alzheimer’s disease, Parkinson’s disease, type 2 diabetes, Creutzfeldt-Jacob
(Mad Cow) disease, and some cancers are related to the presence of
amyloid proteins. Most are incurable
or difficult to treat.
Wide angle view
What Dr Guy has been doing is trying to bring together the vast array of data available to find a hypothesis to explain the possible structure of these smaller polypeptides that are known as oligomers. Experimentally, their shapeshifting ways make detailed molecular structures difficult to determine, but there is a wealth of low resolution data available on their sizes and shapes and effects on neurons plus high resolution data on related fibril structures.

Dr Guy has found one class of structure known as a β-barrel that would be consistent with all of the available information for oligomers and transmembrane channels. Fibrils have multiple layers of β-sheets. The backbone chains of β-strands are stretched as much as possible. These thread-like structures of adjacent monomers bind together to form β-sheets. For smaller assemblies these sheets can roll into cylindrical shapes to form β-barrels. Dr Guy suspects that, like fibrils, these barrels can have multiple layers. The sizes of the barrels can vary enormously and depend upon the length and number of the polypeptide chains that comprise the barrel. In most fibrils, all monomers have identical conformations. If this holds true for the β-barrels it simplifies the modelling process greatly. In fact, radial symmetry can be detected in micrographs of amyloid annular protofibrils and transmembrane channels. Also, β-barrel theory is well developed, so if the number of β-strands and the β-barrel’s diameter and symmetry can be determined the backbone itself is very predictable.
Hypothesis-based research is a cornerstone of contemporary science. Dr Guy’s work has already made suggestions for how certain protein configurations can be stabilised. This may help structural biologists to determine high-resolution structures, neurobiologists to determine which structures are toxic, and pharmaceutical companies to design drugs and develop antibodies that prevent formation of toxic assemblies.
Posing challenges like solving the structure of small oligomers also helps develop our knowledge of the complexities and possibilities of protein folding. This knowledge can be used to refine our understanding of the physical basis of certain diseases. Therefore, knowing why certain protein structures form will also help improve medical diagnostic toolkits and help diagnose and treat people even earlier.
Personal Response
Are there any similar proteins you will be working on next?
The next extension of our modelling could be the most significant. We are modelling how a variety of amyloids bind to Aβ hexamers. These amyloids include synucleins (Parkinson’s disease), amylin (type 2 diabetes), humanin (a mitochondrial micropeptide), and human Cystatin C (a cysteine proteinase inhibitor). Some of these interactions may be detrimental. For example, interactions between Aβ and amylin, may exacerbate both Alzheimer’s disease and diabetes. But other interactions appear to be beneficial; interactions with humanin reduces the toxicity of Aβ. Understanding the structural mechanisms underlying these processes could provide clues for developing new drugs and treatments for these diseases.