X-ray histotomography: Characterising every cell of a whole organism
Diagnosing diseases requires scientists and doctors to understand how cells respond to different medical conditions. Every major disease, from cardiovascular disease and diabetes to cancer, is closely associated with 3-dimensional microscopic cellular and tissue architectural changes that are both typical and informative of disease mechanisms. A common way of studying the microscopic abnormalities of disease is a 2-dimensional method called histology, in which thin slices of centimetre-sized tissue samples are fixed, stained to distinguish cellular components, and examined for abnormal features.
Dr Cheng and his team were searching for a way to study cellular changes digitally and quantitatively, across all cell types in a whole animal model.
This powerful technique revolutionised biology and medicine in the 1800s. Indeed, histology is the foundation of our understanding that all living things are made of cells. It has been used for over a century to visualise cellular composition and tissue architecture in millimetre- to centimetre-scale tissues from diverse multicellular organisms. It is a powerful way to distinguish normal and abnormal cellular features, enabling diagnosis of diseases and important discoveries in biology. Dr Keith Cheng at Pennsylvania State University and his team of biologists, physicists, and engineers were searching for a way to study these changes digitally and quantitatively, across all human diseases and their animal models, in all three physical dimensions – but there was no way to do that.
Overcoming the limits of histology
Histology allows individual cells to be distinguished by cutting tissues into slices less than 1/200th of a millimetre thick. In practice, only a small fraction of any given tissue sample is studied in histology. It is therefore impossible to see the entirety of cells and structures that are thicker than the slice, or to accurately measure 3-dimensional features such as shape or volume. An additional limitation is that physical sectioning is irreversible, making it impossible to recut sections in alternate planes, and physical cutting introduces tissue loss and distortions that compromise our ability to digitally define the 3D structure of tissue and organisms at the cellular level.
To circumvent histology’s limitations, a whole-sample, 3D version of histology is needed to provide us with comprehensive quantitative and volumetric measurements of scientifically and clinically meaningful features. A 3D histology would enable safer drugs, more precise diagnoses, and more in-depth knowledge of the interactive roles of genes, chemicals, and disease in biology.
Micro-computed tomography
Larger internal structures within the human body are routinely visualised using a technique known as computed tomography, CT for short. CT scan imaging is widely used in hospitals. This technique uses X-rays and a computer to create detailed pictures of the inside of the body. It takes X-ray pictures from hundreds to thousands of different angles to derive 3D images computationally. “Micro-CT” works on the same principles as CT scans. While CT scans allow us to see things as small as a millimetre, micro-CT allows us to see things 1000-times smaller – even parts of cells. To achieve 3D histology, both high tissue contrast and high resolution are needed to distinguish and characterise cells and tissues. Since contrast is insufficient in unstained samples, specific metal-based stains are needed for histology-level contrast. The samples for micro-CT can be insects, tissues, vertebrate embryos and zebrafish at any life stage.
Source: https://elifesciences.org/articles/44898/figures
Micro-CT optimisation
Ideally, 3D histology would allow one to rapidly scan for abnormalities in entire organisms and tissue samples and, at the same time, enable the histological examination of any portion of the sample from any angle. No existing method has had the necessary combination of field of view, resolution and contrast. That was, until Dr Cheng and his team adapted lens and camera technologies from the semiconductor and imaging industries. Implementing these improvements would allow scientists and doctors to image even whole small organisms at subcellular resolution.
X-ray histotomography
By optimising micro-CT, Dr Cheng and his team created a 3D histology they called X-ray histotomography as part of a vision to visually and quantitatively compare normal and abnormal microanatomy in 3D, across all living things. X-ray histotomography can achieve the resolution, contrast, and field of view required for histological evaluations and the ability to phenotype all cell types and tissues in whole organisms. This improved form of micro-CT retains the strengths of histology and creates a path to digital histology for full sample volumes.
A primary goal was to implement a technology that can enable large-scale 3D phenotyping. X-ray histotomography was derived from micro-CT by optimising X-ray optics, sample preparation, and digital workflow. The resulting adjustments increased the resolution and potential for high throughput (the speed of data generation and collection). Samples are rigidly embedded in plastic so that they can be preserved for years without deterioration and re-imaged for discovery, validation, and reproducibility. Their system also reduces unwanted sample movement during scanning, resulting in improved image quality. To test the potential of their optimised micro-CT system, the Penn State team needed a model that had the full range of tissue complexity and sample sizes commonly encountered in experimental and clinical histology.
Multiple factors make the zebrafish a useful model for developing whole-organism 3D tissue imaging. Zebrafish measure one millimetre to about a centimetre in width. Indeed, the zebrafish is a well-characterised vertebrate model with diverse tissues, and about 70% of human genes have at least one obvious corresponding gene in zebrafish, explaining its common use as a vertebrate model of human biology and disease.
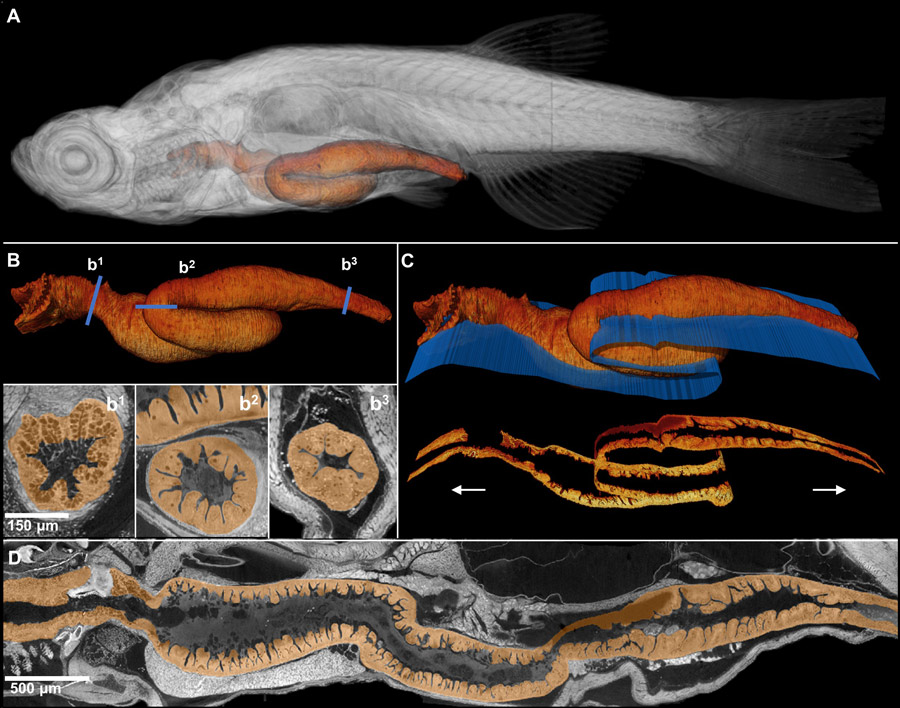
With X-ray histotomography, Dr Cheng and his team were able to obtain 3D digital reconstructions of whole zebrafish. These digital zebrafish allow tissues and cells to be visualised across organ systems in the whole animal. While both traditional histology and histotomography have the resolution needed to distinguish cellular features in 2D slices, only the latter reveals elongated, complex 3D tissue structures such as vessels, nerve tracts, and bones. The digital representation of full volumes of tissue has created an opportunity for an entirely new form of “histology” in which we can digitally “cut” and “re-cut” the tissue at any angle or slice thickness. Parameters can be customised to easily visualise various tissue structures. Structures that are normally inaccessible to histology are accessible when using X-ray histotomography. For example, the team was able to study features of complex structures such as the gills and the gut.
Computational phenotyping for phenomics
The intention of Dr Cheng’s work was to fill a critical gap in developing the transdisciplinary field called phenomics. Phenomics is the study of complete sets of phenotypes across all organisms. Phenotypes are observable traits of organisms that result from interactions between genes, environment, disease, and chance. Any physical feature is a phenotype; for instance, our eye colour is a phenotype that depends on our genes. Nutrition, stress, or temperature and humidity are examples of environmental factors that can impact phenotypes. Every symptom caused by disease is also a phenotype. The purpose of the emerging field of phenomics is to systematically determine, measure, and compare phenotypes across biology to understand the links between specific phenotypes and their causal genetic, environmental, and disease factors.
X-ray histotomography may soon be used to study changes across hundreds of cell types in centimetre-sized organisms or tissue samples.
Phenotyping consists of recording the observed features of an organism. It is of central importance in biology and medicine since phenotyping includes identifying symptoms to make a diagnosis, and to understand sets of mutant phenotypes that illuminate the functions of a gene. Because of the importance of histology, Dr Cheng and his team sought to expand its power in phenotyping in the third dimension. In this way, 3D data of cell and tissue architecture would be used to systematically study how disease, genetics and environmental factors impact organismal phenotypes. 3D microanatomical phenotyping would also improve the accuracy and prognostic implications of diagnoses that can benefit from quantification of histological details. Because phenotypes often affect more than one organ system, the ability of histotomography to detect phenotypes across all cell types makes it ideal for determining phenotypes more completely. Examining every cell type in a whole organism is an ideal way to study the phenotype at both organismal (organ sizes and shapes) and cellular scales. Justifiably, Dr Cheng and a growing team of collaborators are excited about working together towards a whole-organism 3D imaging technique at subcellular resolution and to create scientifically and medically useful computational tools to detect and interpret those changes.
Multiple possibilities and future directions
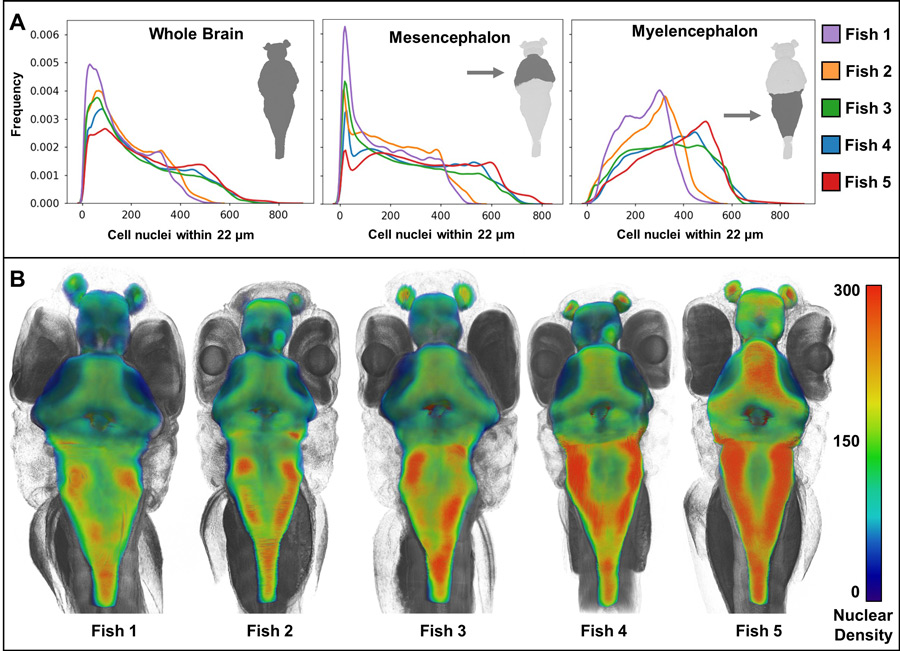
The combination of its 3D nature and centimetre field of view allows X-ray histotomography to enable visualisation of convoluted, millimetre- to centimetre-scale structures such as intestines, bones, blood vessels, and nerve tracts, while retaining the ability to examine cellular change at subcellular resolution. Histological features are usually described qualitatively, but the most useful measures in science are quantitative. X-ray histotomography solves this problem by making it possible to characterise tissue change quantitatively. Histotomography’s unprecedented combination of visualisation of all cell types, high resolution and scale, allow users to analyse cellular features such as size and shape in the context of the whole sample. For example, densities of brain cells were computed and represented in colour maps that revealed striking individual phenotypic variation between zebrafish siblings born on the same day.
Dr Cheng and his team have also confirmed that histotomography can be used to identify pathological phenotypes and make diagnoses. Indeed, minute histopathological features such as nuclear fragmentation have been detected in zebrafish with a hereditary defect in DNA synthesis. The absence of small structures such as the air bladder and its tiny pneumatic duct cannot be confirmed using histology due to their small size and tortuous shape. Because histotomography allows visualisation of the entire organism in exquisite detail, they were able to confirm not only that their mutant never forms those structures, but also that their cells share nuclear features with cancer, such as nuclear atypia.
X-ray histotomography could be used to study changes across hundreds of cell types in any millimetre to centimetre-sized organism or tissue sample. The computational and visual insights into 3D cell and tissue architecture will be useful for reference atlases, comprehensive organismal screens, and diagnostics. Students, teachers, and researchers will also enjoy the fact that histotomography’s 3D tissue and organismal images can be visualised, manipulated, and “cut” using virtual reality technologies.
Current efforts aim at increasing resolution, field of view, and imaging speed, and automating phenotyping and tissue diagnosis. The enabled computational tissue analysis will lead to a better understanding of how genes and the environment contribute to phenotypes, and result in a healthier environment and safer drugs.

Personal Response
What inspired you to believe that optimising micro-CT to obtain cellular resolution was possible?
How will you apply X-ray histotomography in future research?