Epidemiology–laboratory interactions for developing resilience against future infectious diseases
Bacteria are one of the leading causes of public health burden associated with food and waterborne diseases. In addition, they have many abilities that allow them to survive decontamination methods, such as biofilm formation. With the rise in antibiotic resistance and climate change also driving bacterial proliferation, new approaches are needed to mitigate the public health burden of infectious diseases. Professor Aliyar Cyrus Fouladkhah of the Public Health Microbiology Laboratory at Tennessee State University, USA, explores the ecology of bacterial biofilms and antimicrobial interventions against foodborne bacteria. His research group utilises emerging technologies such as application of elevated pressure and novel and plan-based antimicrobials for developing resilience in the food chain against bacterial pathogens of public health concerns.
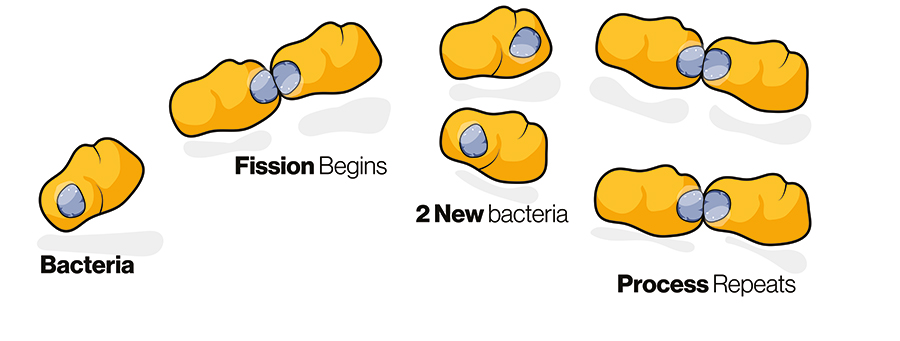
Microbial life – including bacteria, fungi, and viruses – has been present on Earth for around 3.5 billion years, vastly preceding the emergence of humans some 300,000 years ago. Part of the reason these organisms are so successful is due to the way they use their genetic material. Bacteria have a variety of ways in which they can evolve to overcome new challenges and to encourage diversity and fitness, ie, the ability to adapt, survive, and outcompete other strains of bacteria within an environment. One example of these mechanisms is binary fission, which allows one bacterium to multiply into two bacteria – in some cases, this takes as little as 20 minutes. This means that under optimal conditions, a single bacterium is able to multiply to over 50 billion bacteria in the space of 24 hours.
While some bacteria are beneficial, such as probiotic bacteria that are part of the microbial ecosystem in the human gastrointestinal tract, others can have serious consequences for human health and diseases.
Due to their rapid replication rate and their ability to survive in otherwise unhospitable conditions, microbial pathogens present a major public health concern. Already, the World Health Organization estimates that 420,000 individuals die each year due to infections associated with contaminated foods globally, and a further 2 million deaths are due to waterborne diarrhoeal diseases.
Professor Aliyar Fouladkhah, faculty director of the Public Health Microbiology Laboratory at Tennessee State University, studies the ways that foodborne and waterborne bacteria can cause diseases and survive in challenging environments. In understanding the techniques that bacteria use to survive, novel decontamination treatments can be considered to reduce the public health burden of these infectious diseases.
Climate change augments the public health burden of infectious diseases
There are currently two main challenges in the fight against infectious diseases: the rise of antibiotic resistance and the changing climate. Bacteria are constantly finding new ways to push back against existing medications, meaning that when a new therapy is developed, the bacteria might have already started to develop resistance. Researchers must stay one step ahead and consider more novel therapeutic approaches to ensure we have the tools to effectively treat infectious diseases.
Climate change is augmenting the spread and proliferation of microbial pathogens and infectious diseases. Fouladkhah explains that while it can be difficult to predict the impact of a changing climate on infectious diseases, it is well established that increases in environmental temperatures lead to proliferation of bacteria that are able to cause infectious diseases. For example, a 1oC increase in temperature may lead to a 5–10% increase in illnesses caused by Salmonella bacteria (salmonellosis). This could equate to an additional 50,000–100,000 cases of foodborne salmonellosis each year in the USA.
Under optimal conditions, a single bacterium is able to multiply to over 50 billion bacteria in the space of 24 hours.
Fouladkhah highlights the need for new approaches which consider the impact of climate change on antibiotic resistance and infectious diseases. He also explains that it is not just foodborne pathogens that should be considered. Increased rainfall, coupled with changes in environment due to climate change, may contribute to higher multiplication rates of algae and spread of various waterborne infectious diseases.
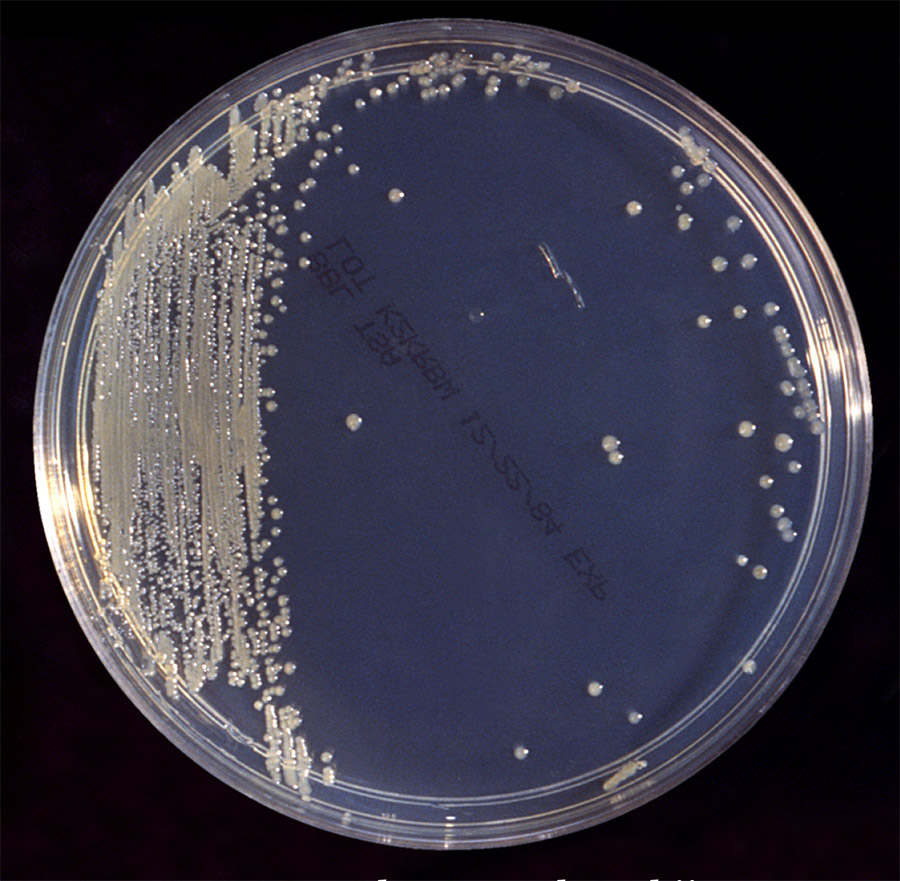
Bacterial biofilms
Bacteria are able to form complex structures or communities called ‘biofilms’, such as the build-up of bacteria on teeth that forms dental plaques. Within these colonies, the bacteria communicate with each other and with bacteria outside the biofilm through a process called quorum sensing. Quorum sensing provides bacteria with the advantage of passing on messages; if one bacterium obtains genetic changes that allow it to become resistant to a drug, it can spread changes in gene regulation throughout the whole colony. This has obvious benefits for bacteria, but it also offers an attractive therapeutic target: disrupting this communication pathway makes bacteria vulnerable to antimicrobial treatments.
Biofilms also help the bacteria to be resistant to some interventions, including the use of antibacterial agents used in food and drink manufacturing. Fouladkhah has previously explored the effectiveness of substances such as sodium hypochlorite against different biofilms. The results of the study showed that sodium hypochlorite was unable to completely eliminate biofilms that were 1–2 weeks old, meaning that decontamination strategies may be less effective than previously thought.
Bacteria such as Cronobacter sakazakii can survive in dry environments, such as powdered infant formula, and can cause fatal infections for premature or newborn infants. Fouladkhah explains that interventions are required to minimise the risk of contamination with the bacteria during the manufacturing of infant formulas. These include existing practices, such as regular sampling and testing for contamination and ensuring that caregivers follow instructions when preparing infant formula, as well as more novel techniques such as the use of high-pressure processing.

Food- and waterborne infectious diseases
Bacterial infections are responsible for the vast majority of foodborne illnesses. The most common pathogens include Escherichia coli, some strains of Salmonella, Listeria monocytogenes, and some species of Cronobacter, as mentioned above. Salmonella alone is the leading cause of deaths caused by foodborne illnesses in the USA, causing around over a million episodes of illness each year.
These bacteria are able to form biofilms, so improper cleaning of surfaces that come into contact with food can be a contributor to foodborne diseases. Fouladkhah compared biofilm formation by E. coli and Salmonella and the response to changes in temperature and exposure to different antimicrobials. The researcher found that increased temperatures caused enhanced biofilm production, and bacteria were more likely to survive after sanitation if they had been exposed to higher temperatures. This may indicate the need for different sanitation protocols for different environmental conditions.
There are currently two main challenges in the fight against infectious diseases: the rise of antibiotic resistance and the changing climate.
In addition to foodborne diseases, bacteria can also cause waterborne diseases. Perhaps the most well-known example is the outbreak of cholera, caused by a bacterium called Vibrio cholerae. Today, waterborne diseases tend to be most commonly associated with many of the same bacteria that cause foodborne illness, such as Salmonella, L. monocytogene, and Shiga toxin-producing E. coli.
In particular, this is due to their ability to survive and develop biofilms. The researchers at Tennessee State University’s Public Health Microbiology laboratory explored this further by investigating how well these bacteria survived in surface water stored at different temperatures (5, 25 and 37oC) and under normal or high-pressure conditions. They found that all three pathogens demonstrated similar survival abilities, with Salmonella having the best chance for survival. This has important implications for public health and food processing as it suggests that these bacteria are able to persist, even in unfavourable and low-nutrient conditions, for extended period of time.

Emerging technologies
Due to their ability to survive many current decontamination approaches, there is a need for new and emerging technologies to reduce the public health burden of food and waterborne infectious diseases. One way this could be achieved is through the use of hydrostatic processing of food products.
Traditional food processing methods that use high temperatures to inactivate microorganisms can impact the quality of the resulting food product. However, the use of non-thermal elevated hydrostatic pressure may offer an attractive alternative. This approach involves applying very high pressures to foods, around 600 MPa or 87 K PSI for under 10 minutes. High-pressure processing techniques are already used for many products, but this approach can be expensive. Therefore, it may be possible to use slightly lower pressures and still see the beneficial effects. A study done by Fouladkhah and colleagues used mild heat and pressure-based pasteurisation methods for milk and orange juice and was able to show that using mild heat alongside pressure-based treatments provided the best results.
Given the rise of antibiotic resistance and the impact of climate change on bacterial survival and proliferation, there is a need to develop new preventative measures. Fouladkhah proposes that this could be done by implementing a combination of epidemiological tools and laboratory technologies, such as genetic sequencing of bacteria. Whole genome sequencing techniques are allowing scientists to quickly detect and characterise outbreaks of infectious diseases. Using new technologies and therapies will help to mitigate some of the impacts of climate change on infectious diseases, but ultimately preventing future outbreaks may be avoided in some part by taking action to reduce our impact on the environment.
Personal Response
What should be the next steps in combatting infectious diseases?
Eradication of polio from the human populations and rinderpest from animal populations in the 20th century are success stories and reminders that we should not lose hope against mitigating the burden of infectious diseases. Adoption and implementation of cutting-edge technologies and analytical methods, development of new innovative and evidence-based therapies and therapeutics, mitigating the deleterious impacts of climate change, and enhanced laboratory and epidemiology interactions will be our best hopes in the future for developing resilience and preventing future outbreaks and sporadic episodes of infectious diseases.